In this post I want to show how to create plots with R (2D and 3D) for different atlases, first. Second, I will demonstrate how plot personal data onto those plots. Therefore I use the data provided by Ganis et al. (2004). As this post serves also as a documentation for a short report1, some details of the paper are mentioned here aswell.
Packages
Atlas packages
Although there are other packages out there (e.g. the 📦 ggbrain-pacakge), we deal with the 📦 ggseg-package because
- it builds on the known Grammar of Graphics, adapted in the 📦
ggplot2-package (Wickham, 2010) - provides everything we need to start (i.e. Desikan-Killany cortical atlas
dk+ Automatic subcortical segmentatioaseg) - with the 📦
ggsegExtra-package it provides a lot of addional atlases (see below) - custom atlases or coloring is possible and
- everything can be ploted as well in 3D via the 📦
ggseg3d-package
Desikan-Killany cortical atlas
The Desikan-Killany cortical atlas dk is provided as a basis by the 📦 ggseg-package. It contains 90 distinct brain regions (to access the regions and information see dk$data).
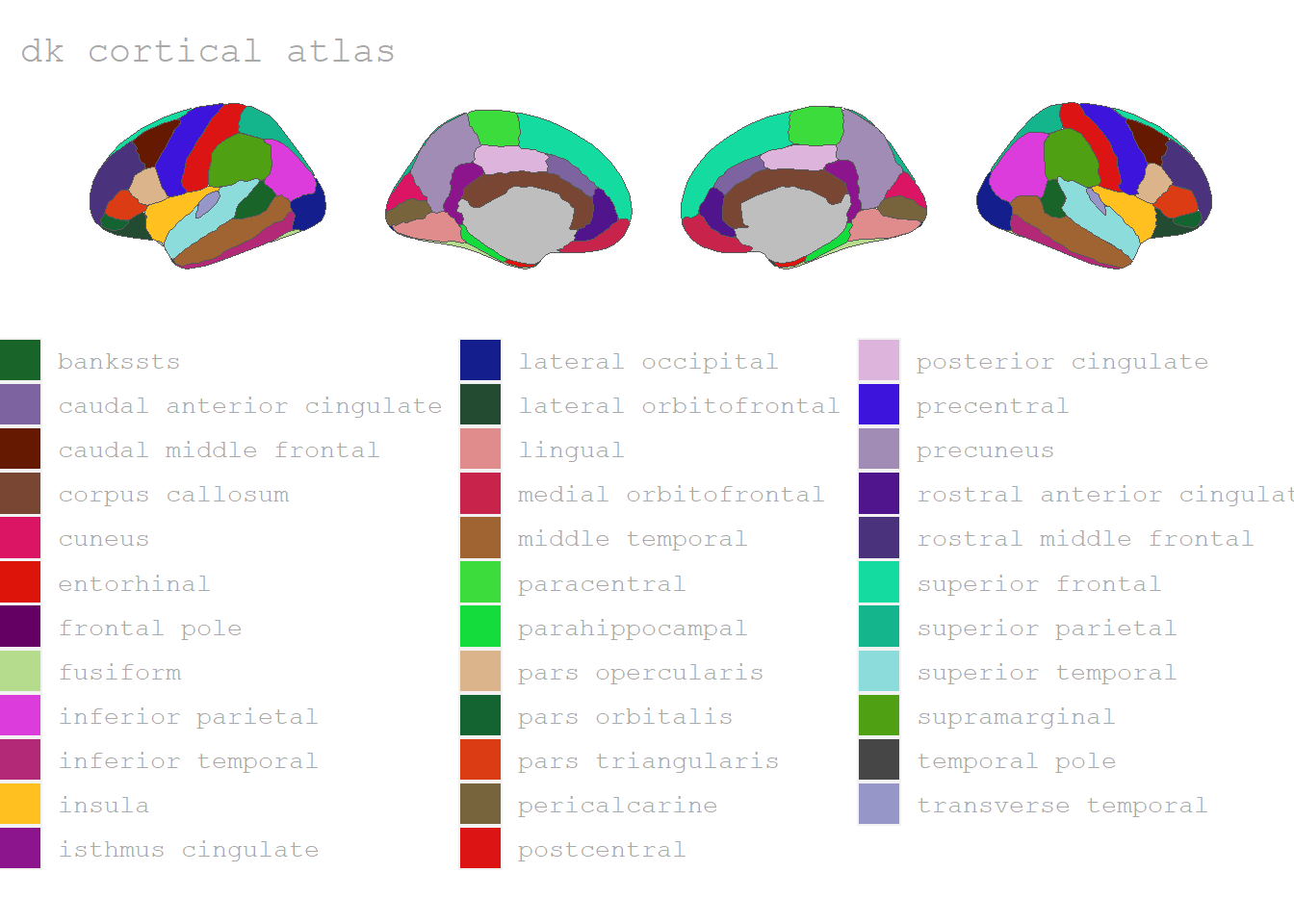
As mentioned, the Desikan-Killany- and all the other cortical atlases can be viewed in 3D as well:
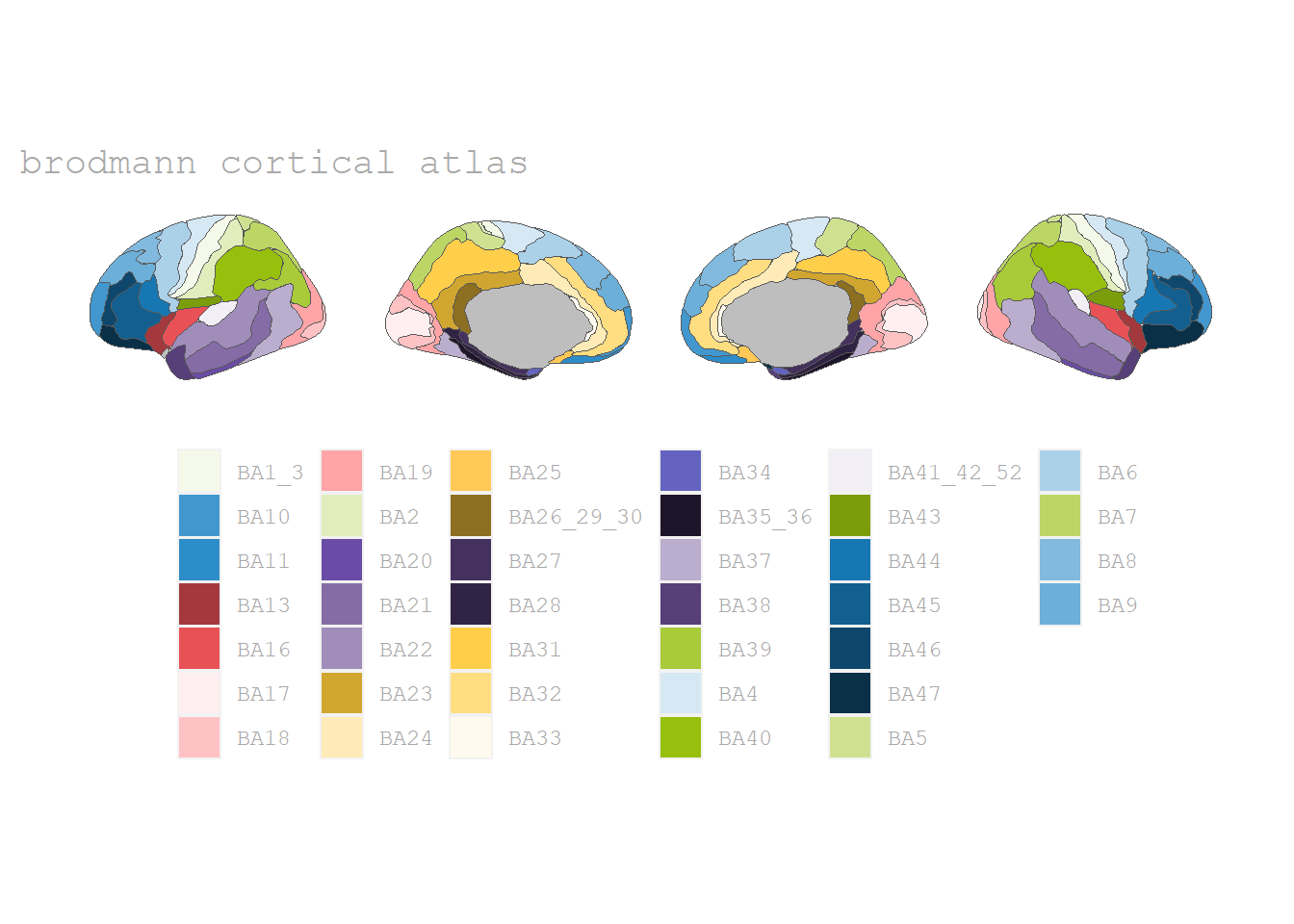
And as said, there are more atlases available, such as the Brodman areas:
Brodman ares
The Brodman ares (Brodman, 1909) are available in the 📦 ggsegBrodmann-package by Pijnenburg et al. (2021).
Flechsig’ atlas of functional segregation
The atlas by Flechsig (1920) is a historical one that contains 46 regions, available via the 📦 ggsegFlechsig-package and also transfered to R by Pijnenburg et al. (2021).
# remotes::install_github("ggseg/ggsegFlechsig")
library(ggsegFlechsig)
library(ggsegExtra)
plot(flechsig)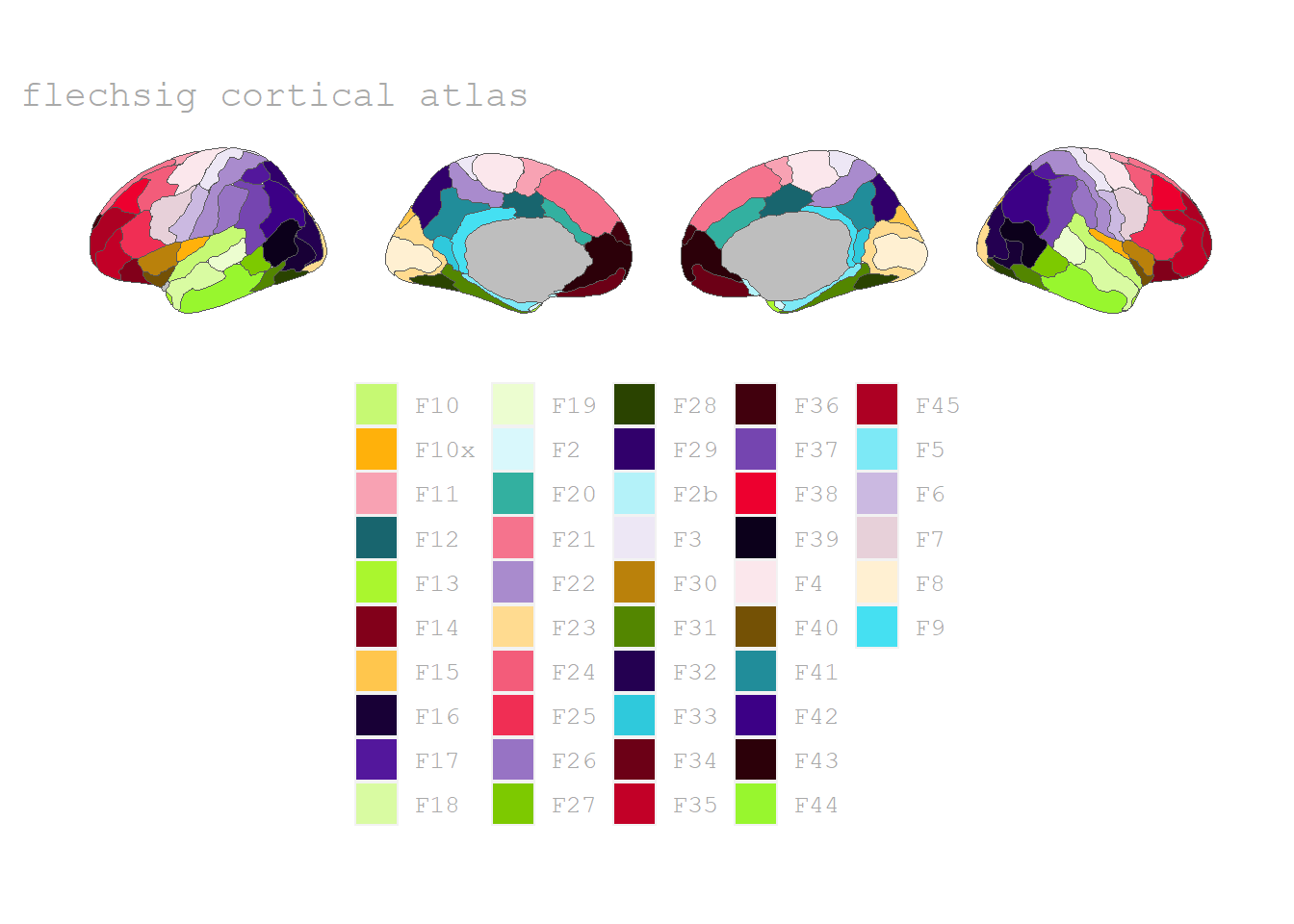
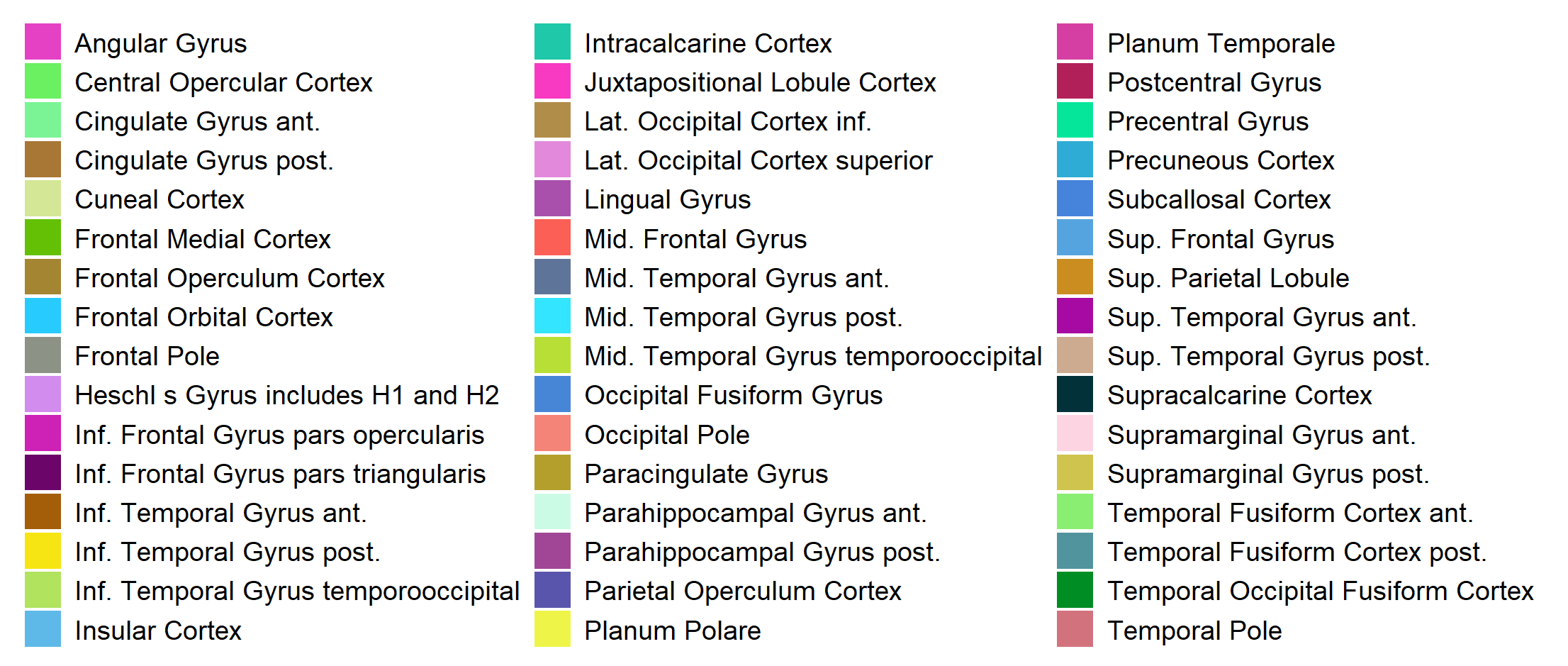
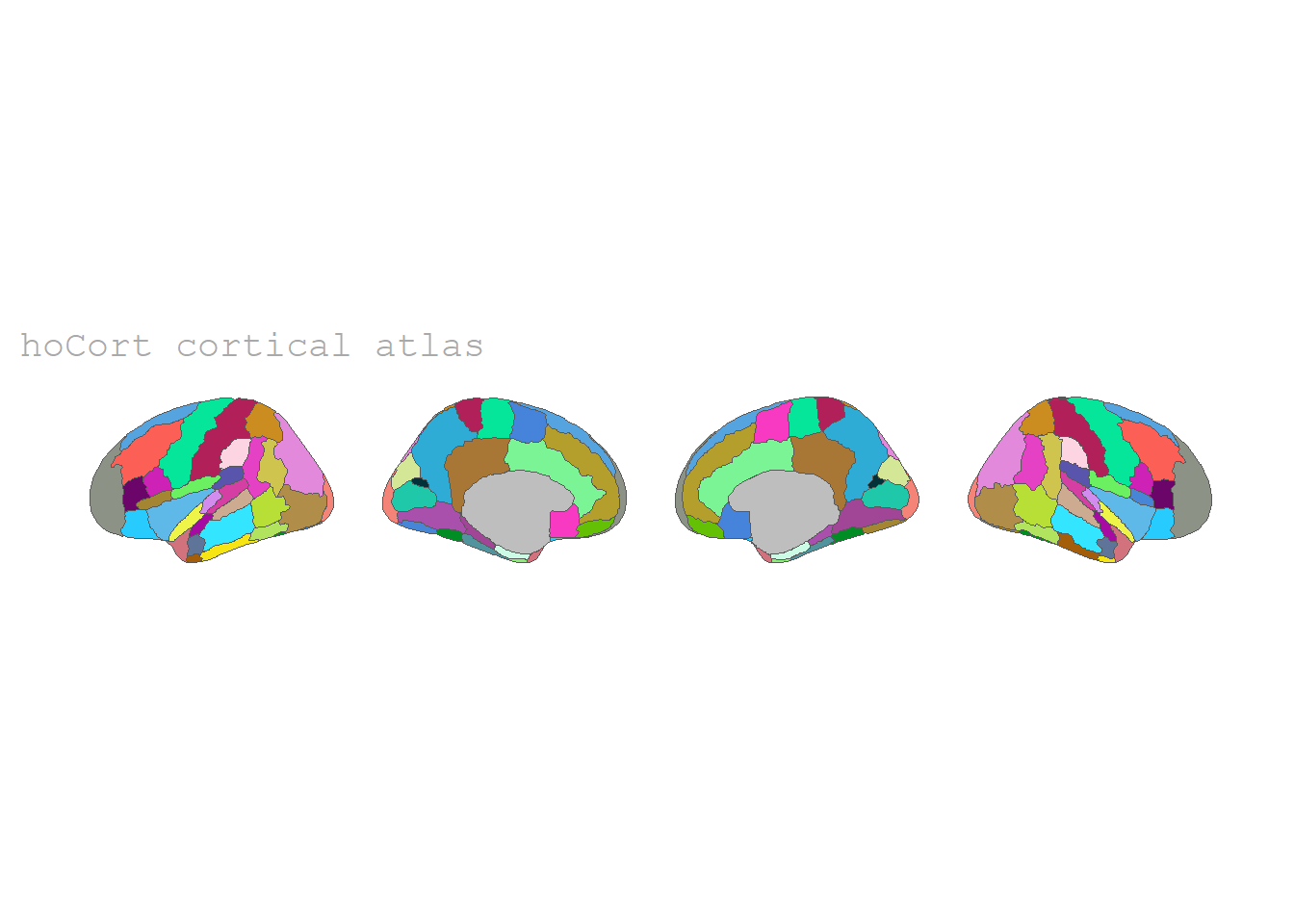
Harvard-Oxford cortical atlas
The Harvard-Oxford cortical atlas is the one we will use in the later example. For more see the paper of Makris et al. (2006).

Example
Paper
To demonstrate how to color a given atlas with external data (e.g. information given by a research paper), I use the data provided by Ganis et al. (2004) in their article Brain areas underlying visual mental imagery and visual perception: an fMRI study.
The studies design can be sumed up as follows: participants had to learn a set of visual stimuli (white objects on black ground, e.g. a tree). Aferwards they underwent both conditions of the experiment: a) Perception (P) and b) Imagery (I) in seperate fMRI scans (counterbalanced).
During the P-condition they saw one of the known images per trail + heard a coresponding sound (e.g. the word tree). An accustic statement given 4.5 s later had to be confirmed (2AFC, e.g. wider than tall coded as W). I-condition was the same execpt subjects had to imagine the image (only sound, no image, eyes closed).
Reaction time data
(Non) responses and accuracy
On each trial, participants pressed one of two keys in response to the probe question. Participants were instructed not to press either key if they could not understand the name at the beginning of a trial because of scanner noise. Overall, the name was missed more often during the imagery than during the perception blocks (20.6% vs. 5.2%, respectively, F(1,14) = 22.32, p < 0.001).
Code
df_resp <- tribble(~"group", ~"h_reaction_no",
"imagery", 0.206,
"perception", 0.052) %>%
mutate(h_reaction_yes = 1-h_reaction_no)
theme_set(theme_minimal())
df_resp %>% pivot_longer(cols = starts_with("h_reaction"),
names_to = "reaction",
names_prefix = "h_reaction_",
values_to = "h") %>%
mutate(label = paste0(round(h*100, 1), "%")) %>%
ggplot(aes(x = "", y = h, fill = reaction)) +
geom_col(alpha = .8) +
geom_label(aes(label = label),
position = position_stack(vjust = 0.5),
# size = 8,
show.legend = FALSE) +
scale_fill_manual(values = c("lightcoral", "steelblue3")) +
coord_polar(theta = "y") +
facet_wrap(~group) +
theme(# text = element_text(size = 30),
axis.title = element_blank(),
axis.text = element_blank()) 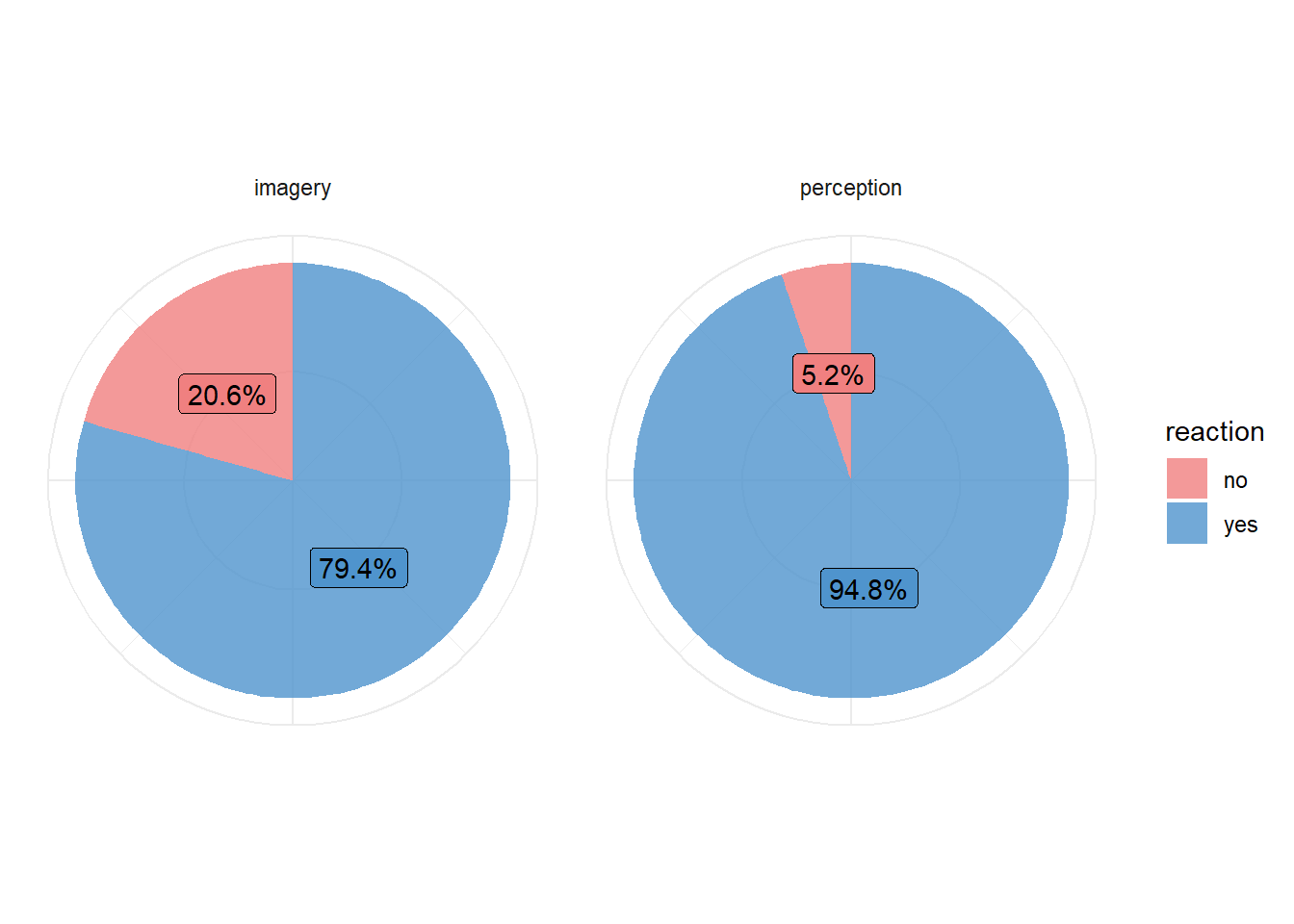
However, for the trials on which a response was made, accuracy was very high and did not differ between imagery and perception (96.2% vs. 97.3% correct, respectively, F(1,14) = 0.78, p>0.1)
Code
df_resp_acc <- tribble(~"condition", ~"h_reaction_correct",
"imagery", 0.962,
"perception", 0.973) %>%
mutate(h_reaction_incorrect = 1-h_reaction_correct) %>%
pivot_longer(cols = starts_with("h_reaction"),
names_to = "reaction",
names_prefix = "h_reaction_",
values_to = "h") %>%
mutate(label = paste0(round(h*100, 1), "%"))
df_resp_acc %>% ggplot(aes(x = reaction, y = h)) +
geom_bar(aes(fill = reaction),
show.legend = FALSE, alpha = .75,
stat = "identity", position = "dodge") +
scale_y_continuous("", labels = scales::label_percent()) +
# scale_fill_viridis_d(option = "G", direction = -1) +
scale_fill_manual(values = c("limegreen", "red")) +
facet_wrap(~condition, scales = "free_x") +
# theme(text = element_text(size = 30)) +
geom_label(aes(label = label),
alpha = .85,
# size = 8,
position = position_stack(vjust = 0.5),
show.legend = FALSE)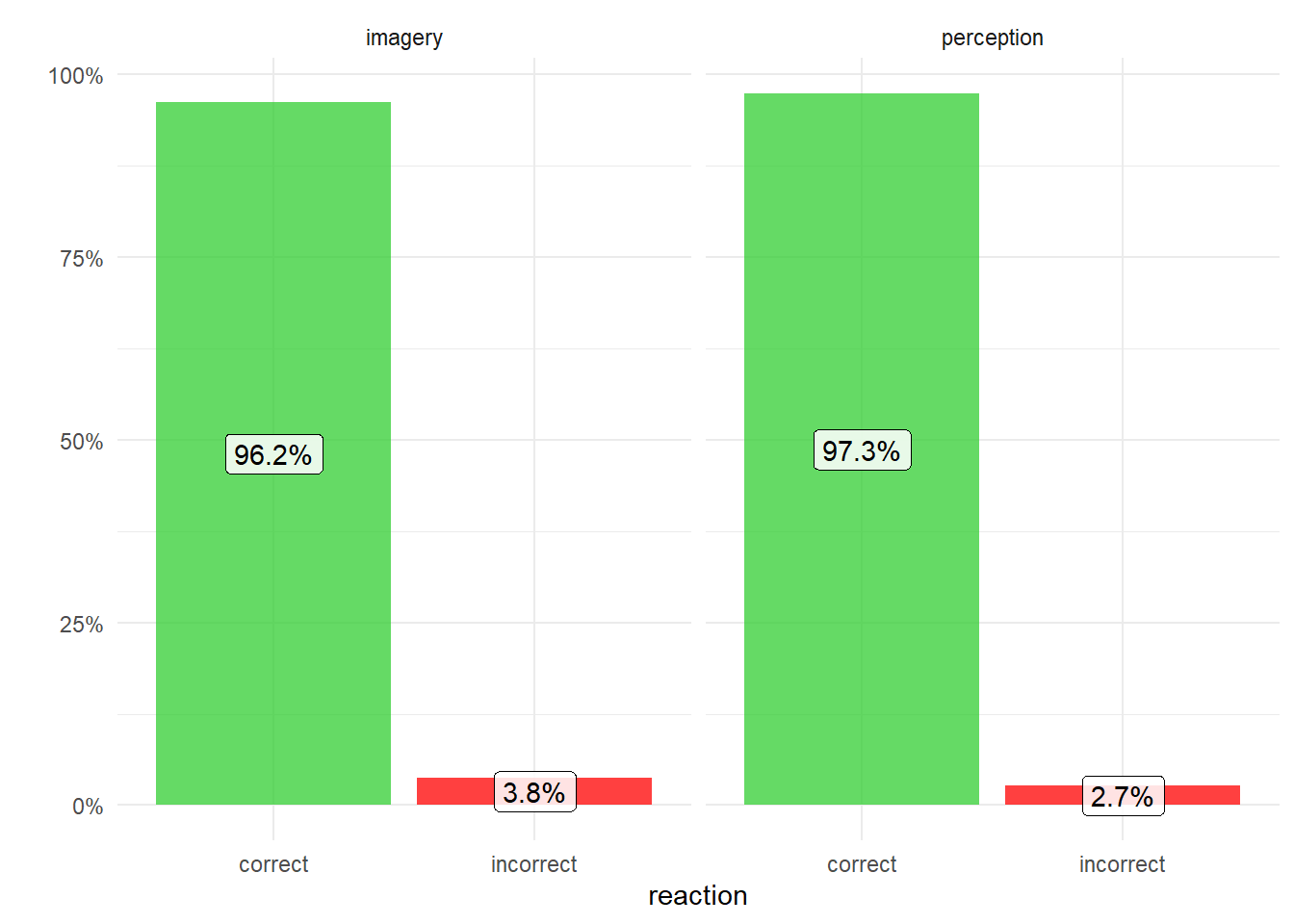
Reaction times
The response times (RTs) for correct trials were slower in the imagery condition than in the perception condition (medians: 1384 vs. 1232 ms, respectively, F(1,14) = 6.3, p = 0.025).
The same analysis was performed on a subgroup of 12 (out of 15) participants, excluding the 3 participants with the highest Imagery/Perception RT ratios. For this subgroup, there were no reliable differences between the median RTs in the two conditions (1299 vs. 1208 ms, respectively, F(1,14) = 2, p = 0.18)
Code
# tribble(~ "group", ~".y.", ~"group1", ~"group2", ~"p", ~"p.adj.signif", ~"y.position",
# 1, "RT", "imagery", "perception", .025, "*", 1400,
# 2, "RT", "imagery", "perception", .18, "ns.", 1320) -> stat.test
tribble(~"group", ~"condition", ~"RT", # Mdn
1, "imagery", 1384,
1, "perception", 1232,
2, "imagery", 1299,
2, "perception", 1208) -> df_mdn_rts
df_mdn_rts %>% mutate(group = factor(group, levels = c(1,2),
labels = c("sample (n = 15)",
"subsample (n = 12)"))) %>%
ggplot(aes(x = condition, y = RT)) +
geom_col(aes(fill = condition), show.legend = FALSE) +
labs(y = "Mdn RT [ms]") +
coord_cartesian(ylim = c(1100, 1500)) +
facet_wrap(~ group) +
# ggpubr::stat_pvalue_manual(stat.test, label = "p", tip.length = 0.01)
# theme(text = element_text(size = 30)) +
scale_fill_viridis_d()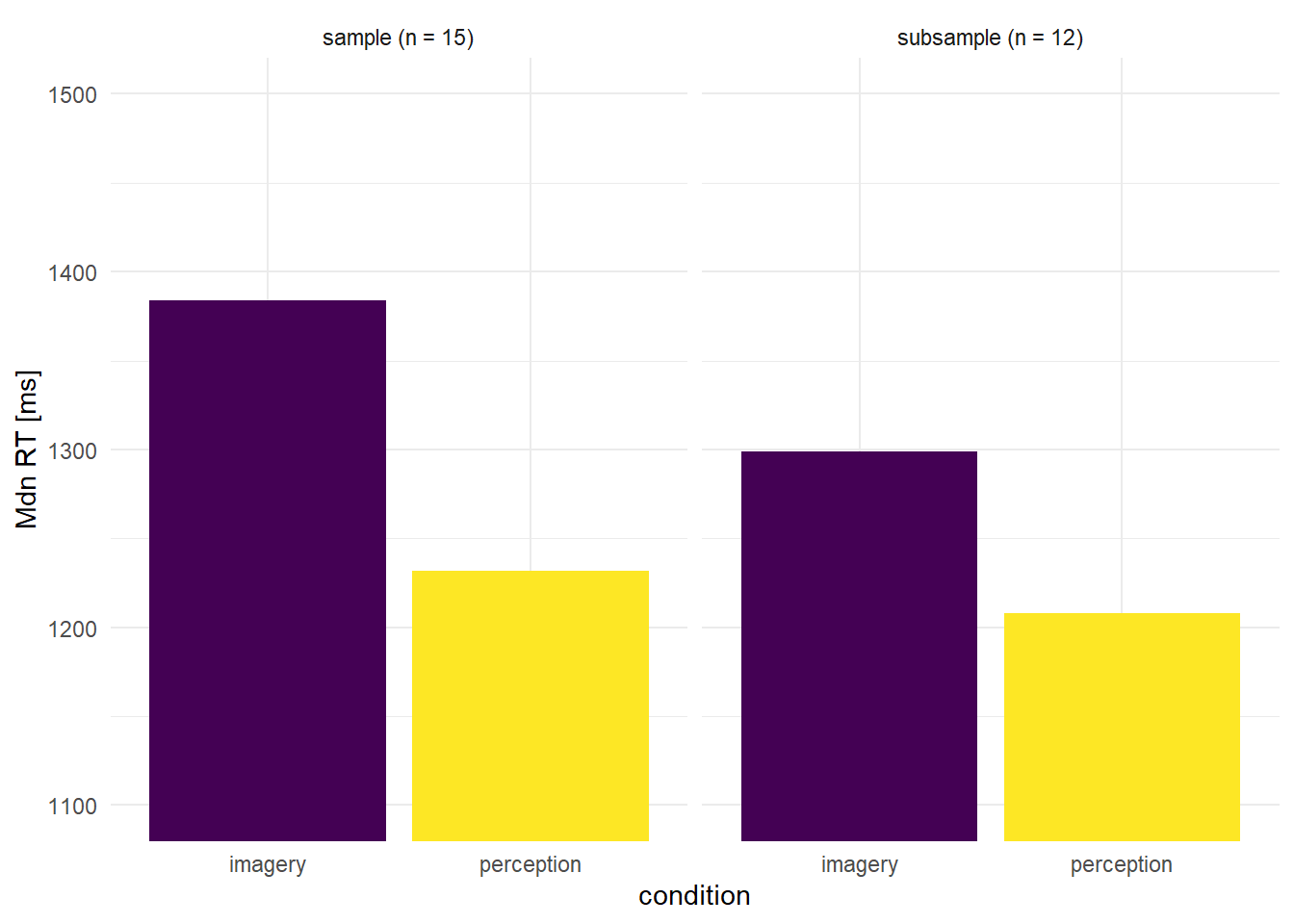
fMRI Data
Quantification of overlap
As the title already suggests, the brain regions that are activated during visual perception are compared to those activated during imagination. The authors quantify the amount of overlap (measured in voxels) as:
- number of common voxels (C): voxels for which the contrast Perception vs. Imagery was n.s. and that exhibited a sig. activation change (rel. to baseline) during both imagery and perception
- number of different voxels (D): voxels for which the contrast Perception vs. Imagery was significant and that exhibited a significant activation (rel. to baseline), in at least one condition
- percentage of shared voxels (S): defined as:
\[S=100\times\frac{C}{C+D}\]
However, for all three quanfifications there are datasets (tables in the paper). I will deal with them in the following order2:
-
Common voxels (
tab_3) -
Different voxels (
tab_4) and -
Shared voxels (
tab_2)
The tables in the paper are too long to be displayed on a PowerPoint slide in any meaningful way. I have therefore tried to transform the tables from long- to wide-shape. Nevertheless, you still can’t get any information from the tables. Therefore the tables are shown as bar-charts, where an activation in the corresponding areas is shown in color. The same colors are then used to color the corresponding brain areas. This should make it possible to see more quickly during the presentation which areas are relevant.
As said, all three of those datasets are included in this blog, however it is not neccessary to read the code for all three of them, so you may skip those parts and just look at the the results (i.e. shared voxels).
Possible contrasts
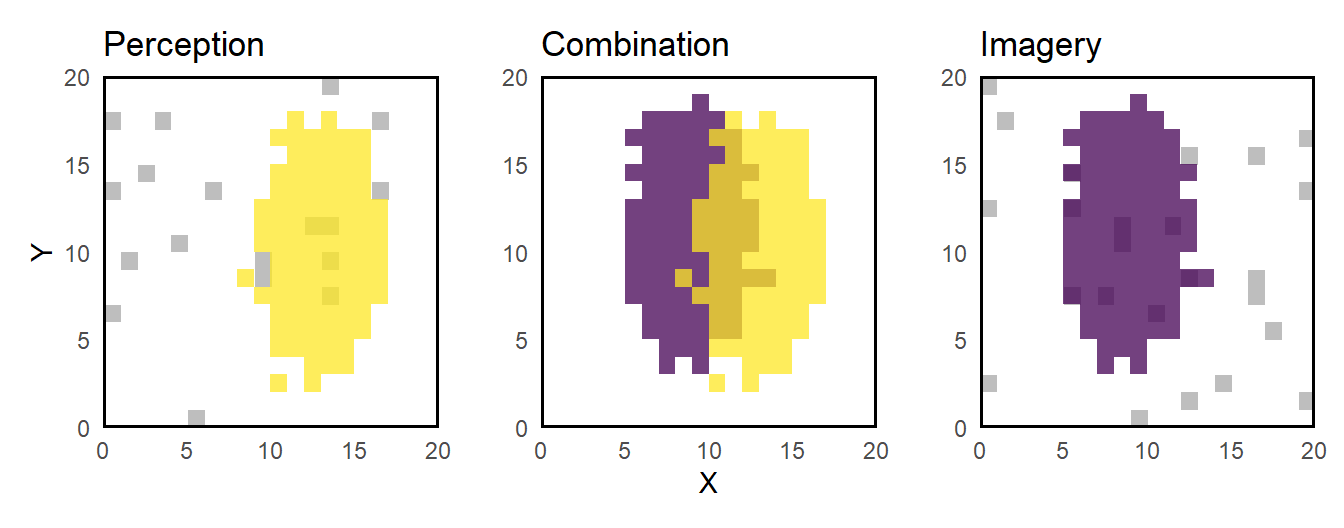
The authors only report significant contrasts (p = .0001). That’s why it’s important to understand which contrasts can exist in the first place. In visualization the gray patern symbolises the baseline activation, both activations (I and P) are sig. different from that, they are partly common and partly exclusive (what does not mean different).
- S: Voxels are referred to as shared if they occur in any of the following conditions (C and D). They are computed according to the formula above, no differentiation is made with respect to any contrasts.
- C: Voxels are referred to as common if a sig. activation change (rel. to baseline) is shown during both conditions, I and P, (i.e. [P>0, I>0] and [P<0, I<0]) and the same voxcels are activ in both conditions (i.e. [P-I=0]). Hence, there are only two possible contrasts: [P>0, I>0, P-I=0] and [P<0, I<0, P-I=0]
- D: Voxels are referred to as different if a sig. activation change (rel. to baseline) is shown during at least one condition (i.e. [P>0, I=0], [P<0, I=0], [P=0, I>0], [P=0, I<0], [P>0, I>0], [P<0, I<0] but not [P<0, I>0] or [P>0, I<0]). Additionally, the contrast P vs I had to be significant (i.e. [P-I>0] or [P-I<0]). Hence, there are 12 possible contrasts:
\[ \small \begin{bmatrix} (P>0, I=0, P−I>0), & (P=0, I>0, P−I>0), & (P>0, I>0, P−I>0) \\ (P>0, I=0, P−I<0), & (P=0, I>0, P−I<0), & (P>0, I>0, P−I<0) \\ (P<0, I=0, P−I>0), & (P=0, I<0, P−I>0), & (P<0, I<0, P−I>0) \\ (P<0, I=0, P−I<0), & (P=0, I<0, P−I<0), & (P<0, I<0, P−I<0) \end{bmatrix} \]
Note. Significant differences are visible in only three of the shown 12 possible contrasts, i.e. (P>0, I=0, P−I>0), (P<0, I=0, P−I<0) and (P>0, I>0, P−I>0).
Reference dataframe
For beeing able to create a custom plot of the brain with the data provided above we have to make sure the names of the areas match with the names of the regions provided in the atlas (dk). Therefore we create a seperate dataframe. For the case when our data matched more then one reagion in the Desikan-Killiany Cortical Atlas we include two (or as many colums) as needed, e.g. we have data for the Middle frontal gyrus but since dk distinguished between rostral and middle frontal gyrus, we create two colums for that. For the other way arround (the atlas provides only one region for several colums of our data) we would choose a different atlas for the ones provided.
Harvard-Oxford cortical atlas Dict
Code
tribble(~"no", ~"region", ~"reg_abrv",
1, "Angular Gyrus", "AG",
2, "Central Opercular Cortex", NA,
3, "Frontal Operculum Cortex", NA,
4, "Frontal Orbital Cortex", NA,
5, "Frontal Pole", NA,
6, "Heschl s Gyrus includes H1 and H2 ", "TransTG",
7, "Inf. Frontal Gyrus pars opercularis", "IFG",
8, "Inf. Frontal Gyrus pars triangularis", "IFG",
9, "Inf. Temporal Gyrus ant.", "ITG",
10, "Inf. Temporal Gyrus post.", "MOG",
11, "Inf. Temporal Gyrus temporooccipital", "MOG", # ~
12, "Insular Cortex", "Insula",
13, "Lat. Occipital Cortex inf.", "IOG",
14, "Lat. Occipital Cortex superior", "SOG",
15, "Mid. Frontal Gyrus", "MFG",
16, "Mid. Temporal Gyrus ant.", "MTG",
17, "Mid. Temporal Gyrus post.", "MTG",
18, "Mid. Temporal Gyrus temporooccipital", "MTG", #~
19, "Occipital Fusiform Gyrus", "FG",
20, "Occipital Pole", "MOG", #~
21, "Parietal Operculum Cortex", "IPL", # not good
22, "Planum Polare", NA,
23, "Planum Temporale", NA,
24, "Postcentral Gyrus", "postCG",
25, "Precentral Gyrus", "PreCG",
26, "Sup. Frontal Gyrus", "SFG",
27, "Sup. Parietal Lobule", "SPL",
28, "Sup. Temporal Gyrus ant.", "STG",
29, "Sup. Temporal Gyrus post.", "STG",
30, "Supramarginal Gyrus ant.", "SMG",
31, "Supramarginal Gyrus post.", "SMG",
32, "Temporal Occipital Fusiform Cortex", "FG",
33, "Temporal Pole", NA,
34, "Cingulate Gyrus ant.", "AntCingG",
35, "Cingulate Gyrus post.", "postCingG",
36, "Cuneal Cortex", "Cun",
37, "Frontal Medial Cortex", "MeFG",
38, "Intracalcarine Cortex", NA,
39, "Juxtapositional Lobule Cortex", NA,
40, "Lingual Gyrus", "lingG",
41, "NA", NA,
42, "Paracingulate Gyrus", "CingG", # ?
43, "Parahippocampal Gyrus ant.", "paraHG",
44, "Parahippocampal Gyrus post.", "paraHG",
45, "Precuneous Cortex", "preCuneus",
46, "Subcallosal Cortex", NA,
47, "Supracalcarine Cortex", NA,
48, "Temporal Fusiform Cortex ant.", "FG",
49, "Temporal Fusiform Cortex post.", "FG"
) %>% select(-no) -> df_regio_abr2
#' nicht abgetragen:
#' Cerebellum - Cereb
#' LentNuc
#' Caudate
#' Claustrum
#' Thalamus
#'
#' tab_3 %>% select(brain_region) %>% data.frame() %>% distinct() %>% filter(!brain_region %in% c("lingG", "Insula", "TransTG", "AG", "Cun", "postCingG", "AntCingG", "paraHG", "preCuneus", "postCG", "PreCG", "FG", "SFG", "IFG", "MFG", "MeFG", "SOG", "ITG", "MTG", "STG", "MOG", "SPL", "SMG", "CingG", "IPL", "IOG"))Desikan-Killiany Cortical Atlas Dict (old)
Code
# old version
tribble(~"reg_abrv", ~"reg_name", ~"persp", ~"region", # region refers to dk region
# FRONTAL
"IFG", "Inferior frontal gyrus", "lateral", "pars orbitalis",
"IFG", "Inferior frontal gyrus", "lateral", "pars opercularis",
"IFG", "Inferior frontal gyrus", "lateral", "pars triangularis",
"MFG", "Middle frontal gyrus", "lateral", "caudal middle frontal",
"MFG", "Middle frontal gyrus", "lateral", "rostral middle frontal",
"SFG", "Superior frontal gyrus", "lateral", "superior frontal",
"PreCG", "Precentral gyrus", "lateral", "precentral",
# PARIETAL
"postCG", "Postcentral gyrus", "lateral", "postcentral",
"IPL", "Inferior parietal lobule", "lateral", "inferior parietal",
"SPL", "Superior parietal lobule", "lateral", "superior parietal",
"AG", "Angular gyrus", "subcort", NA, # not good! lateral but not on map
"SMG", "Supramarginal gurus", "lateral", "supramarginal",
# TEMPORAL
"ITG", "Inferior temporal gyrus", "lateral", "inferior temporal",
"MTG", "Middle temporal gyrus", "lateral", "middle temporal",
"STG", "Superior temporal gyrus", "lateral", "superior temporal",
"STG", "Superior temporal gyrus", "lateral", "bankssts", # ~
"TransTG", "Transverse temporal gyrus", "lateral", "transverse temporal",
# = Heschl’sche Querwindungen
# OCCIPITAL
"SOG", "Superior occipital gyrus", "lateral", NA, #
"MOG", "Middle occipital gyrus", "lateral", "lateral occipital",
"MOG", "Middle occipital gyrus", "lateral", "pericalcarine",
"IOG", "Inferior occipital gyrus", "lateral", NA, # ???
"MeFG", "Medial frontal gyrus", "medial", "paracentral", #??
# evtl. nur sup
"Insula", "Insula", "lateral", "insula",
"AntCingG", "Anterior cingulate gyrus", "medial", "caudal anterior cingulate",
"AntCingG", "Anterior cingulate gyrus", "medial", "rostral anterior cingulate",
"postCingG", "Posterior cingulate gyrus", "medial", "isthmus cingulate",
"paraHG", "Parahippocampal gyrus", "medial", "parahippocampal",
"FG", "Fusiform gyrus", "medial", "fusiform",
"lingG", "Lingual gyrus", "medial", "lingual",
"preCuneus", "Precuneus", "medial", "precuneus",
"Cun", "Cuneus", "medial", "cuneus",
"LentNuc", "Lenticular nucleus", "subcort", NA,
# Ncl. lentiformis = Putamen + Globus pallidus
"Claustrum", "Claustrum", "subcort", NA,
"Thalamus", "Thalamus", "subcort", NA,
"Caudate", "Caudate nucleus", "subcort", NA,
"CingG", "Cingulate gyrus", "medial", "posterior cingulate", # not sure what they mean since cing is present above
"Cereb", "Cerebellum", "subcort", NA) -> df_regio_abr
# not sure if it's Cerebellum but can't be Cerebrum
# cf.:
# dk$data %>% data.frame() %>% select(region) %>% filter(!is.na(region)) %>% filter(grepl('dings', region))Common voxels (C)
Altough the data is provided by a published article, there are some mistakes in the table for which we have to take account for, otherwise the plots won’t be correct. So after reading in the data we have to clean it (as shown below):
- areas that have got different names in the other (upcoming) tables (e.g. ling vs. lingG should be the same)
- areas that are named differently within the same datatable (e.g. Claustrum is once spelled Claustrun and since displayed as two differnt areas in the barcharts below)
- additional problems, e.g. left and right postCG appears two times each in the table for shared voxels, what might be a) a mistake or b) because of missing talairach coordinates (x, y, z; cf. datatable 3 [below] and 4). We remove the doubled values since we do not know whether a or b is the case.
- additional problem is caused not by the paper but its (Latex) format, i.e. the minus sign that is coded as a long minus (“—”) aka. “\002”.
Code
tab_3 <- read_table(here("data", "neuro", "tab_3.txt"),
col_types = list(side = col_character(),
brain_region = col_character(),
perc_overlap = col_double(),
C_voxels = col_character(),
x = col_character(),
y = col_character(),
z = col_character())) %>%
mutate(side = factor(side, levels = c("L", "R"), labels = c("left", "right")),
brain_region = if_else(brain_region == "Claustrun", "Claustrum",
brain_region),
C_voxels = stringr::str_replace_all(C_voxels, "\\*|\\(|\\)", "")) %>%
mutate_at(.vars = c("x", "y", "z"),
~stringr::str_replace_all(., "\\002", "-")) %>%
mutate_at(.vars = c("x", "y", "z"),
~stringr::str_replace_all(., ",", "")) %>%
mutate_at(.vars = c("C_voxels", "x", "y", "z"), as.numeric) # %>%
# distinct() # no! Values are distinct, cf. contrasts
tab_3 %<>% mutate(contrast = c(rep(1, 60), rep(2, 13))) %>%
mutate(contrast = factor(contrast, levels = c(1, 2),
labels = c("P>0, I>0", "P<0, I<0"))) %>%
distinct()Plots
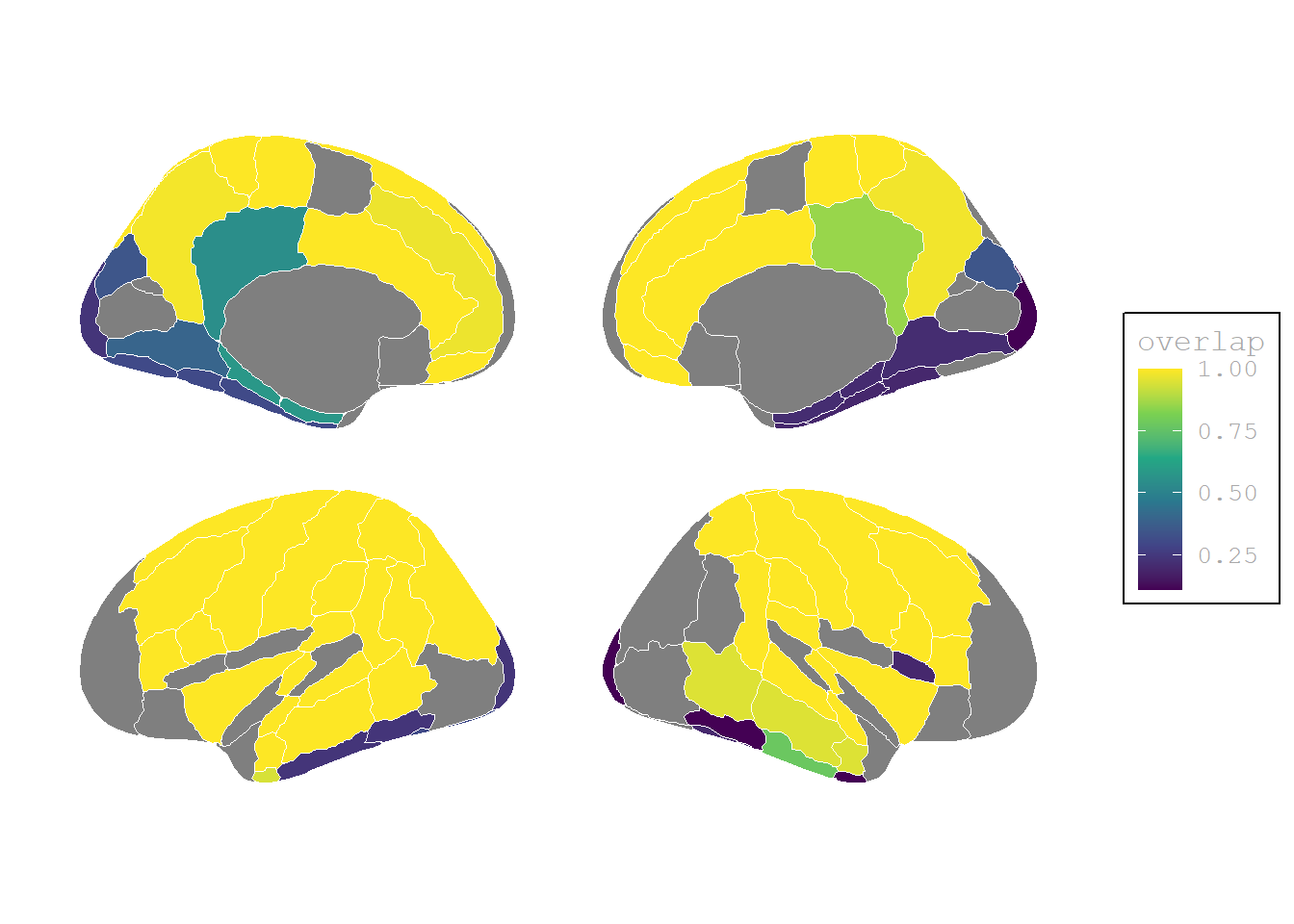
Activation maps
(P>0, I>0) Activation stronger than baseline
Code
theme_set(theme_brain2())
tab_3 %>% filter(contrast == "P>0, I>0") %>%
# left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
left_join(df_regio_abr2, by = c("brain_region" = "reg_abrv")) %>%
rename(hemi = side) %>%
# mutate_at(scale, .vars = vars(C_voxels)) %>%
mutate(overlap = perc_overlap/100) %>%
select(-c(brain_region, # reg_name,
perc_overlap)) %>%
filter(!is.na(region)) %>%
ggplot() +
scale_fill_viridis_c(option = "D", direction = 1) +
geom_brain(atlas = hoCort,
# atlas = dk,
color = "white",
position = position_brain(side ~ hemi),
mapping = aes(fill = overlap)) +
labs(fill = "overlap") +
theme(# legend.position = "none",
plot.background = element_blank())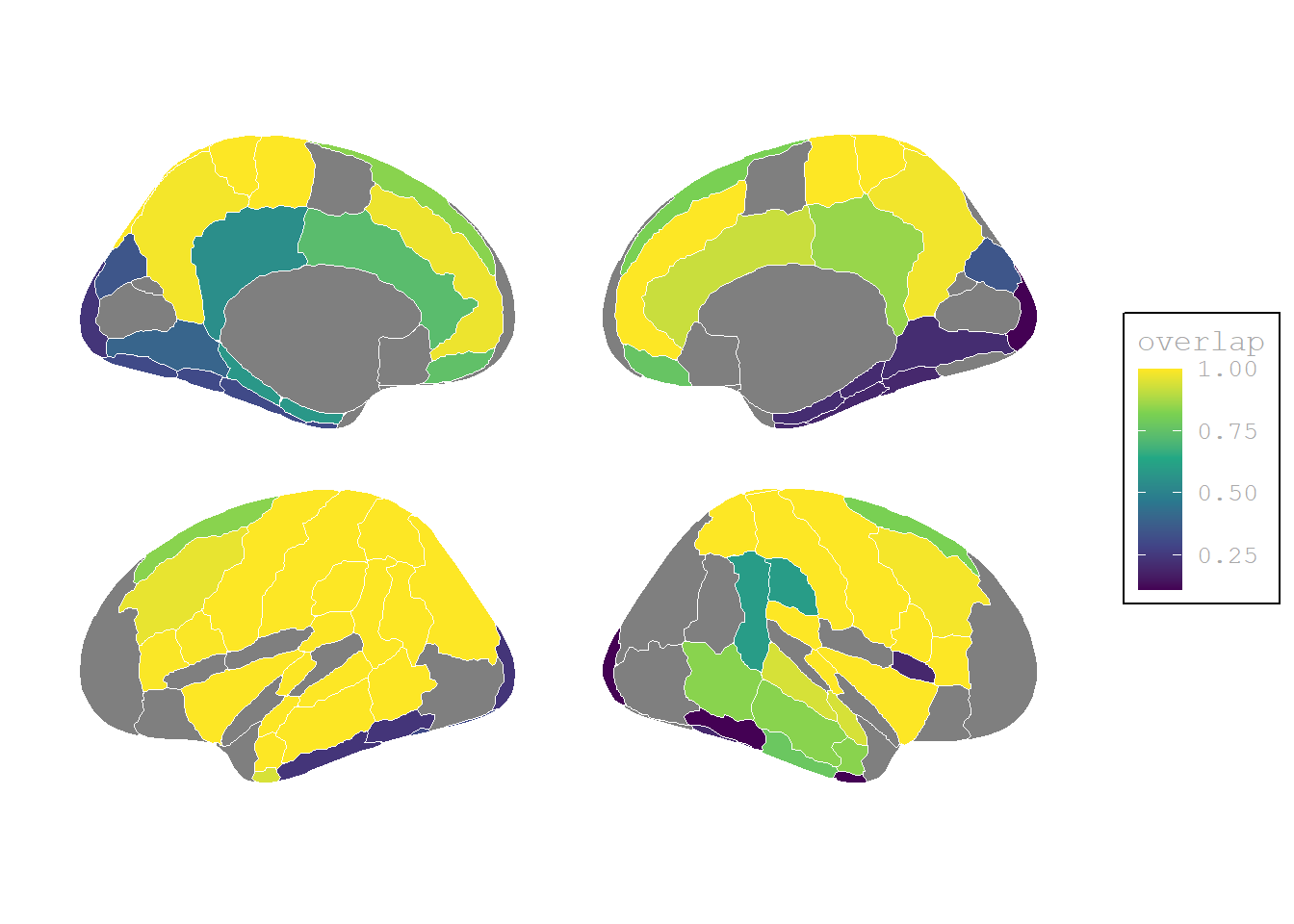
(P<0, I<0) Activation weaker than baseline
Code
theme_set(theme_brain2())
tab_3 %>% filter(contrast == "P<0, I<0") %>%
# left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
left_join(df_regio_abr2, by = c("brain_region" = "reg_abrv")) %>%
rename(hemi = side) %>%
# mutate_at(scale, .vars = vars(C_voxels)) %>%
mutate(overlap = perc_overlap/100) %>%
select(-c(brain_region, # reg_name,
perc_overlap)) %>%
filter(!is.na(region)) %>%
ggplot() +
scale_fill_viridis_c(option = "D", direction = 1) +
geom_brain(atlas = hoCort, # atlas = dk,
color = "white",
position = position_brain(side ~ hemi),
mapping = aes(fill = overlap)) +
labs(fill = "overlap") +
theme(# legend.position = "none",
plot.background = element_blank())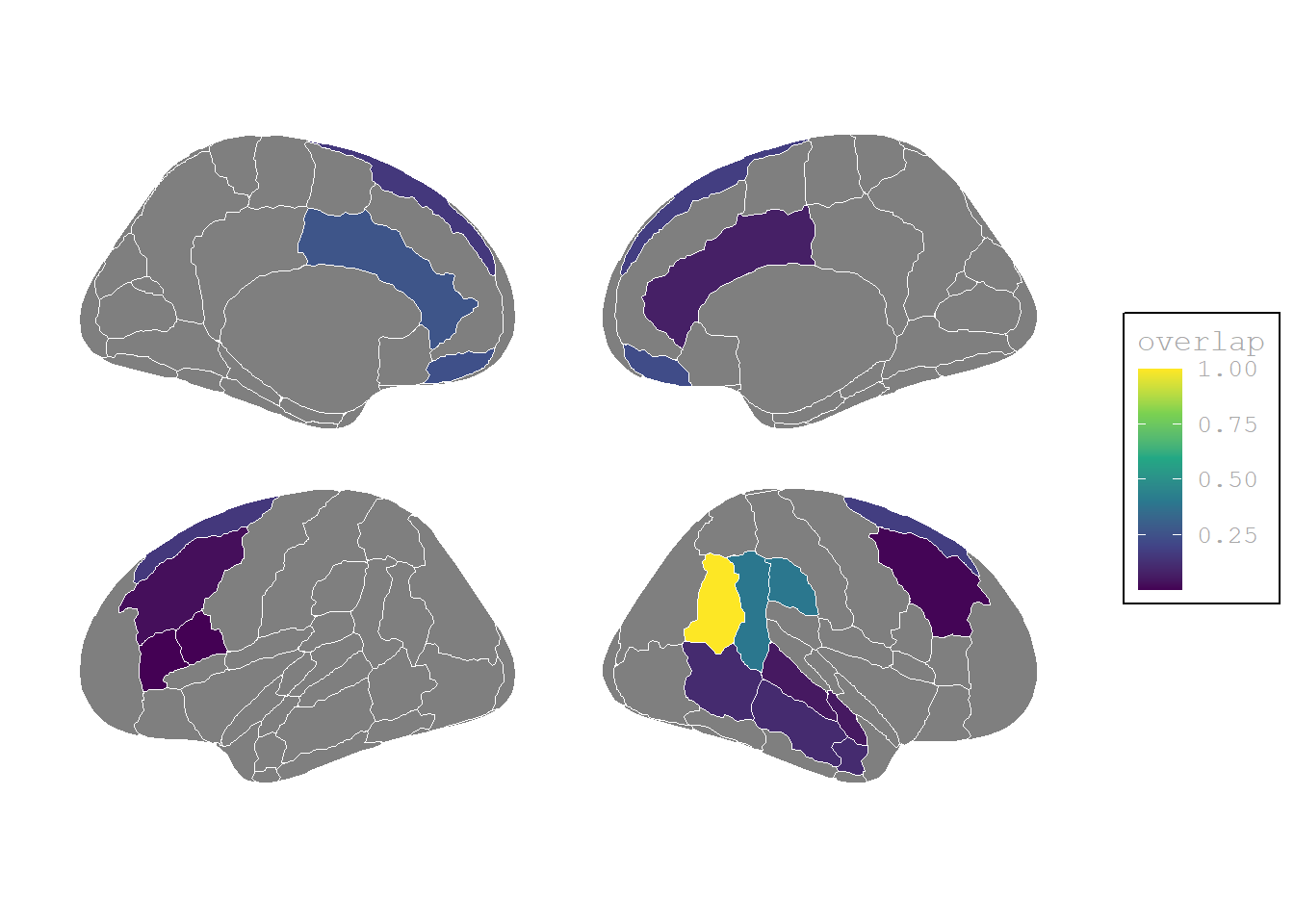
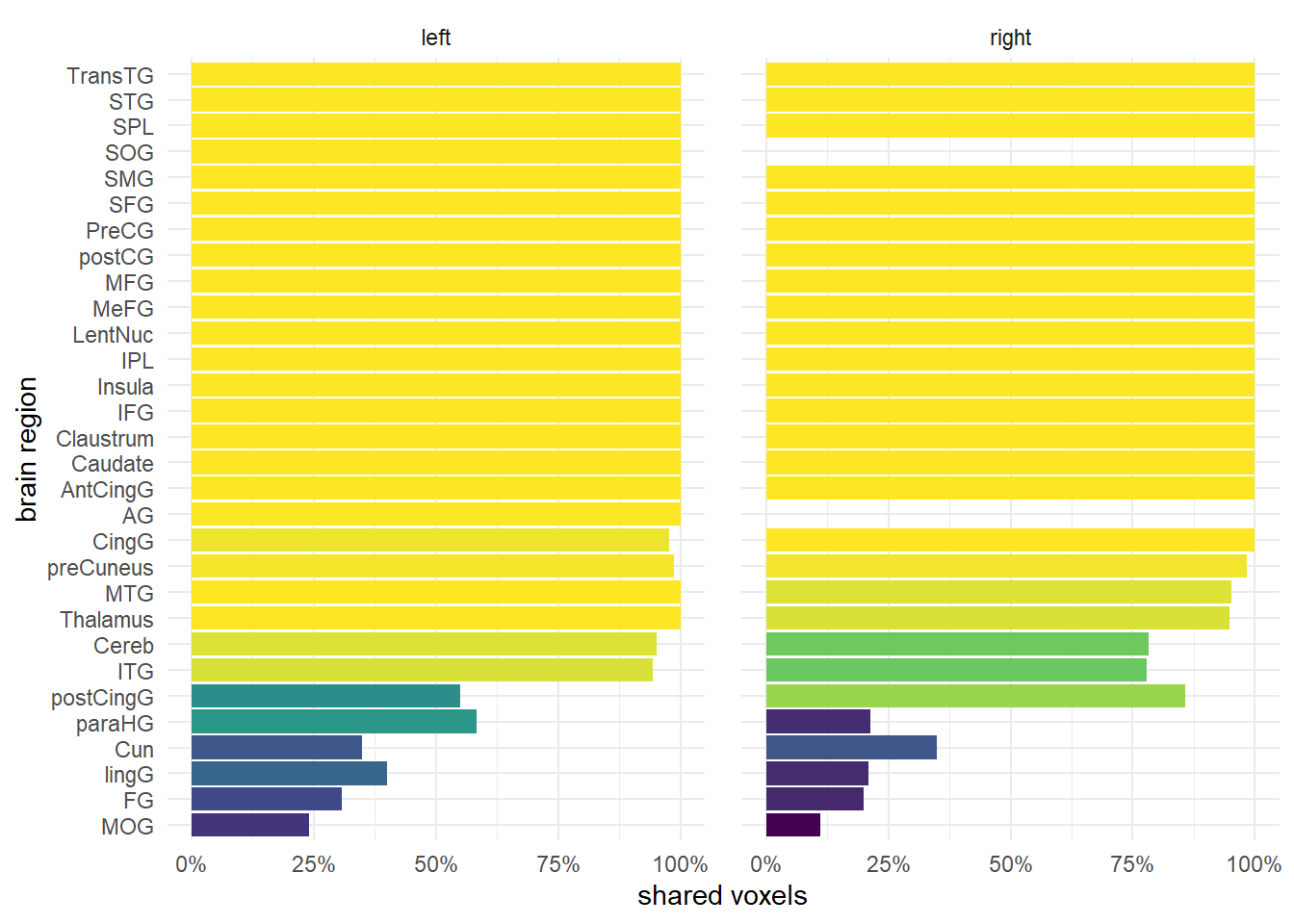
Activation barchart
Only significant contrasts are shown (p = 0.0001), i.e. activation was a) significant higher than at the baseline (P>0, I>0) or significant lower (P<0, I<0) than the baseline for both conditions, but the activation patters were not significant different than zero, i.e. they were similar (P-I=0)
Code
theme_set(theme_minimal())
# df_regio_abr %>% select(reg_abrv, persp) %>%
# distinct() %>%
# right_join(tab_3, by = c("reg_abrv" = "brain_region")) %>%
# # filter(persp != "subcort") %>%
# mutate(rel_overlap = perc_overlap/100) %>%
# # tab_3 %>% mutate(rel_overlap = perc_overlap/100) %>%
# ggplot(aes(x = reorder(reg_abrv, rel_overlap),
# y = rel_overlap, fill = rel_overlap)) +
# geom_bar(stat = "identity") +
# scale_fill_viridis_c(option = "viridis", guide = "none", direction = 1) +
# scale_y_continuous("common voxels", labels = scales::label_percent()) +
# facet_grid(contrast~side, scales = "free", space = "free", drop = TRUE) +
# # theme(axis.ticks.y = element_blank()) +
# labs(x = "brain region") +
# theme(text = element_text(size = 20),
# panel.spacing = unit(0.5, "cm")) +
# coord_flip()
# # theme_minimal()
tab_3 %>% mutate(rel_overlap = perc_overlap/100) %>%
ggplot(aes(x = reorder(brain_region, rel_overlap),
y = rel_overlap, fill = rel_overlap)) +
geom_bar(stat = "identity") +
scale_fill_viridis_c(option = "viridis", guide = "none", direction = 1) +
scale_y_continuous("common voxels", labels = scales::label_percent()) +
facet_grid(contrast~side, scales = "free", space = "free", drop = TRUE) +
# theme(axis.ticks.y = element_blank()) +
labs(x = "brain region") +
theme(# text = element_text(size = 20),
panel.spacing = unit(0.5, "cm")) +
coord_flip() -> vxl_chrt_c
vxl_chrt_c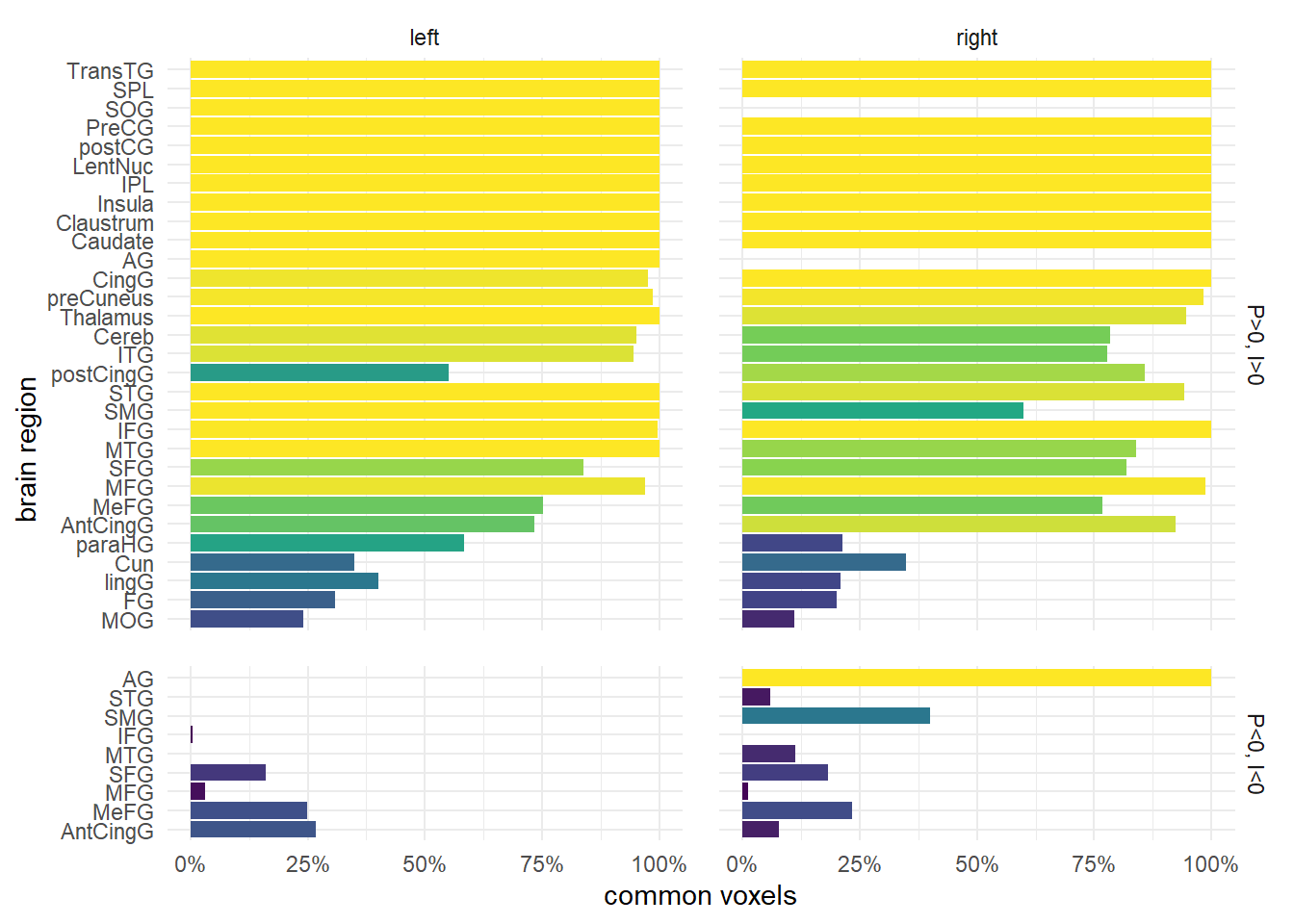
Table C voxels
long to wide tab
Code
tab_3 %>% pivot_wider(names_from = "side",
values_from = c("perc_overlap", "C_voxels", "x", "y", "z")) %>%
relocate("C_voxels_left", .before = "perc_overlap_left") %>%
relocate("C_voxels_right", .before = "perc_overlap_right") %>%
relocate("y_left", .after = "x_left") %>%
relocate("z_left", .after = "y_left") -> tab_3_wide
df_regio_abr %>% select(c(reg_abrv, reg_name, persp)) %>%
distinct() %>%
right_join(tab_3_wide, by = c("reg_abrv" = "brain_region")) %>%
mutate(brain_region = paste0(reg_name, " (", reg_abrv, ")")) %>%
# filter(persp != "lateral") %>%
select(-c(reg_name, persp, reg_abrv)) %>%
# tab_3_wide %>% left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
# mutate(brain_region = paste0(reg_name, " (", brain_region, ")")) %>%
# select(-c(reg_name, persp)) %>%
gt(rowname_col = "brain_region",
groupname_col = "contrast") %>%
tab_stubhead(label = "Contrast") %>%
tab_spanner(label = "left",
columns = c(C_voxels_left, perc_overlap_left, x_left, y_left, z_left)) %>%
tab_spanner(label = "right",
columns = c(C_voxels_right, perc_overlap_right, x_right, y_right, z_right)) %>%
# tab_spanner(label = "left",
# columns = c(x_left, y_left, z_left)) %>%
# tab_spanner(label = "right",
# columns = c(x_right, y_right, z_right)) %>%
# cols_merge(columns = c(S_voxels_left, perc_overlap_left),
# pattern = "{1} ({2})") %>%
# cols_merge(columns = c(S_voxels_right, perc_overlap_right),
# pattern = "{1} ({2})") %>%
# cols_label(S_voxels_left = "S voxels (%)",
# S_voxels_right = "S voxels (%)")
cols_label(C_voxels_left = "C voxels",
perc_overlap_left = "%",
C_voxels_right = "C voxels",
perc_overlap_right = "%",
x_left = "x", y_left = "y", z_left = "z",
x_right = "x", y_right = "y", z_right = "z") %>%
sub_missing() %>%
cols_align(align = "center", columns = everything()) %>%
as_raw_html()| Contrast | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| C voxels | % | x | y | z | C voxels | % | x | y | z | |
| P>0, I>0 | ||||||||||
| Inferior frontal gyrus (IFG) | ||||||||||
| Middle frontal gyrus (MFG) | ||||||||||
| Superior frontal gyrus (SFG) | ||||||||||
| Precentral gyrus (PreCG) | ||||||||||
| Postcentral gyrus (postCG) | ||||||||||
| Inferior parietal lobule (IPL) | ||||||||||
| Superior parietal lobule (SPL) | ||||||||||
| Angular gyrus (AG) | ||||||||||
| Supramarginal gurus (SMG) | ||||||||||
| Inferior temporal gyrus (ITG) | ||||||||||
| Middle temporal gyrus (MTG) | ||||||||||
| Superior temporal gyrus (STG) | ||||||||||
| Transverse temporal gyrus (TransTG) | ||||||||||
| Superior occipital gyrus (SOG) | ||||||||||
| Middle occipital gyrus (MOG) | ||||||||||
| Medial frontal gyrus (MeFG) | ||||||||||
| Insula (Insula) | ||||||||||
| Anterior cingulate gyrus (AntCingG) | ||||||||||
| Posterior cingulate gyrus (postCingG) | ||||||||||
| Parahippocampal gyrus (paraHG) | ||||||||||
| Fusiform gyrus (FG) | ||||||||||
| Lingual gyrus (lingG) | ||||||||||
| Precuneus (preCuneus) | ||||||||||
| Cuneus (Cun) | ||||||||||
| Lenticular nucleus (LentNuc) | ||||||||||
| Claustrum (Claustrum) | ||||||||||
| Thalamus (Thalamus) | ||||||||||
| Caudate nucleus (Caudate) | ||||||||||
| Cingulate gyrus (CingG) | ||||||||||
| Cerebellum (Cereb) | ||||||||||
| P<0, I<0 | ||||||||||
| Inferior frontal gyrus (IFG) | ||||||||||
| Middle frontal gyrus (MFG) | ||||||||||
| Superior frontal gyrus (SFG) | ||||||||||
| Angular gyrus (AG) | ||||||||||
| Supramarginal gurus (SMG) | ||||||||||
| Middle temporal gyrus (MTG) | ||||||||||
| Superior temporal gyrus (STG) | ||||||||||
| Medial frontal gyrus (MeFG) | ||||||||||
| Anterior cingulate gyrus (AntCingG) | ||||||||||
contrasts side by side
Code
df_regio_abr %>% select(c(reg_abrv, reg_name, persp)) %>%
distinct() %>%
right_join(tab_3, by = c("reg_abrv" = "brain_region")) %>%
mutate(brain_region = paste0(reg_name, " (", reg_abrv, ")"),
contrast = recode_factor(contrast, "P>0, I>0" = "A",
"P<0, I<0" = "B")) %>%
# labels = c("P>0, I>0", "P<0, I<0")))
# filter(persp != "lateral") %>%
select(-c(reg_name, persp, reg_abrv)) %>%
pivot_wider(values_from = c(perc_overlap, C_voxels, x, y, z),
names_from = "contrast", values_fill = NA) %>%
gt(rowname_col = "brain_region",
groupname_col = "side") %>%
tab_stubhead(label = "Brain region") %>%
tab_spanner(label = "P>0, I>0",
columns = ends_with("_A")) %>%
tab_spanner(label = "P<0, I<0",
columns = ends_with("_B")) %>%
cols_label(C_voxels_A = "C voxels",
perc_overlap_A = "%",
C_voxels_B = "C voxels",
perc_overlap_B = "%",
x_A = "x", y_A = "y", z_A = "z",
x_B = "x", y_B = "y", z_B = "z") %>%
sub_missing() %>%
cols_align(align = "center", columns = everything()) %>%
as_raw_html()| Brain region | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| % | C voxels | x | y | z | % | C voxels | x | y | z | |
| left | ||||||||||
| Inferior frontal gyrus (IFG) | ||||||||||
| Middle frontal gyrus (MFG) | ||||||||||
| Superior frontal gyrus (SFG) | ||||||||||
| Precentral gyrus (PreCG) | ||||||||||
| Postcentral gyrus (postCG) | ||||||||||
| Inferior parietal lobule (IPL) | ||||||||||
| Superior parietal lobule (SPL) | ||||||||||
| Angular gyrus (AG) | ||||||||||
| Supramarginal gurus (SMG) | ||||||||||
| Inferior temporal gyrus (ITG) | ||||||||||
| Middle temporal gyrus (MTG) | ||||||||||
| Superior temporal gyrus (STG) | ||||||||||
| Transverse temporal gyrus (TransTG) | ||||||||||
| Superior occipital gyrus (SOG) | ||||||||||
| Middle occipital gyrus (MOG) | ||||||||||
| Medial frontal gyrus (MeFG) | ||||||||||
| Insula (Insula) | ||||||||||
| Anterior cingulate gyrus (AntCingG) | ||||||||||
| Posterior cingulate gyrus (postCingG) | ||||||||||
| Parahippocampal gyrus (paraHG) | ||||||||||
| Fusiform gyrus (FG) | ||||||||||
| Lingual gyrus (lingG) | ||||||||||
| Precuneus (preCuneus) | ||||||||||
| Cuneus (Cun) | ||||||||||
| Lenticular nucleus (LentNuc) | ||||||||||
| Claustrum (Claustrum) | ||||||||||
| Thalamus (Thalamus) | ||||||||||
| Caudate nucleus (Caudate) | ||||||||||
| Cingulate gyrus (CingG) | ||||||||||
| Cerebellum (Cereb) | ||||||||||
| right | ||||||||||
| Inferior frontal gyrus (IFG) | ||||||||||
| Middle frontal gyrus (MFG) | ||||||||||
| Superior frontal gyrus (SFG) | ||||||||||
| Precentral gyrus (PreCG) | ||||||||||
| Postcentral gyrus (postCG) | ||||||||||
| Inferior parietal lobule (IPL) | ||||||||||
| Superior parietal lobule (SPL) | ||||||||||
| Angular gyrus (AG) | ||||||||||
| Supramarginal gurus (SMG) | ||||||||||
| Inferior temporal gyrus (ITG) | ||||||||||
| Middle temporal gyrus (MTG) | ||||||||||
| Superior temporal gyrus (STG) | ||||||||||
| Transverse temporal gyrus (TransTG) | ||||||||||
| Middle occipital gyrus (MOG) | ||||||||||
| Medial frontal gyrus (MeFG) | ||||||||||
| Insula (Insula) | ||||||||||
| Anterior cingulate gyrus (AntCingG) | ||||||||||
| Posterior cingulate gyrus (postCingG) | ||||||||||
| Parahippocampal gyrus (paraHG) | ||||||||||
| Fusiform gyrus (FG) | ||||||||||
| Lingual gyrus (lingG) | ||||||||||
| Precuneus (preCuneus) | ||||||||||
| Cuneus (Cun) | ||||||||||
| Lenticular nucleus (LentNuc) | ||||||||||
| Claustrum (Claustrum) | ||||||||||
| Thalamus (Thalamus) | ||||||||||
| Caudate nucleus (Caudate) | ||||||||||
| Cingulate gyrus (CingG) | ||||||||||
| Cerebellum (Cereb) | ||||||||||
Different voxels (D)
Code
tab_4 <- read_table(here("data", "neuro", "tab_4.txt"),
col_types = list(side = col_character(),
brain_region = col_character(),
perc_overlap = col_double(),
D_voxels = col_character(),
x = col_character(),
y = col_character(),
z = col_character())) %>%
mutate(side = factor(side, levels = c("L", "R"),
labels = c("left", "right")),
brain_region = if_else(brain_region == "Claustrun",
"Claustrum", brain_region),
D_voxels = stringr::str_replace_all(D_voxels, "\\*|\\(|\\)", "")) %>%
mutate_at(.vars = c("x", "y", "z"),
~stringr::str_replace_all(., "\\002", "-")) %>%
mutate_at(.vars = c("x", "y", "z"),
~stringr::str_replace_all(., ",", "")) %>%
mutate_at(.vars = c("D_voxels", "x", "y", "z"), as.numeric) # %>%
# distinct() # no! Values are distinct, cf. contrasts
tab_4 %<>% mutate(contrast = c(rep(1, 3), rep(2, 19), rep(3, 4))) %>%
mutate(contrast = factor(contrast, levels = c(1:3),
labels = c("...,I>0,...", "P>0, I=0, P-I>0", "P<0,...,P-I<0")))Plots
Activation maps
The authors report only three contrasts with significantly different activation, i.e. (P>0, I=0, P−I>0), (P<0, I=0, P−I<0) and (P>0, I>0, P−I>0). For the last two of them this is only true for few areas:
(P>0, I=0, P−I>0)
Code
theme_set(theme_brain2())
tab_4 %>% filter(contrast == "P>0, I=0, P-I>0") %>%
# left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
left_join(df_regio_abr2, by = c("brain_region" = "reg_abrv")) %>%
rename(hemi = side) %>%
# mutate_at(scale, .vars = vars(C_voxels)) %>%
mutate(overlap = perc_overlap/100) %>%
select(-c(brain_region, # reg_name,
perc_overlap)) %>%
filter(!is.na(region)) %>%
ggplot() +
scale_fill_viridis_c(option = "D", direction = 1) +
geom_brain(atlas = hoCort, # atlas = dk,
color = "white",
position = position_brain(side ~ hemi),
mapping = aes(fill = overlap)) +
labs(fill = "overlap") +
theme(# legend.position = "none",
plot.background = element_blank())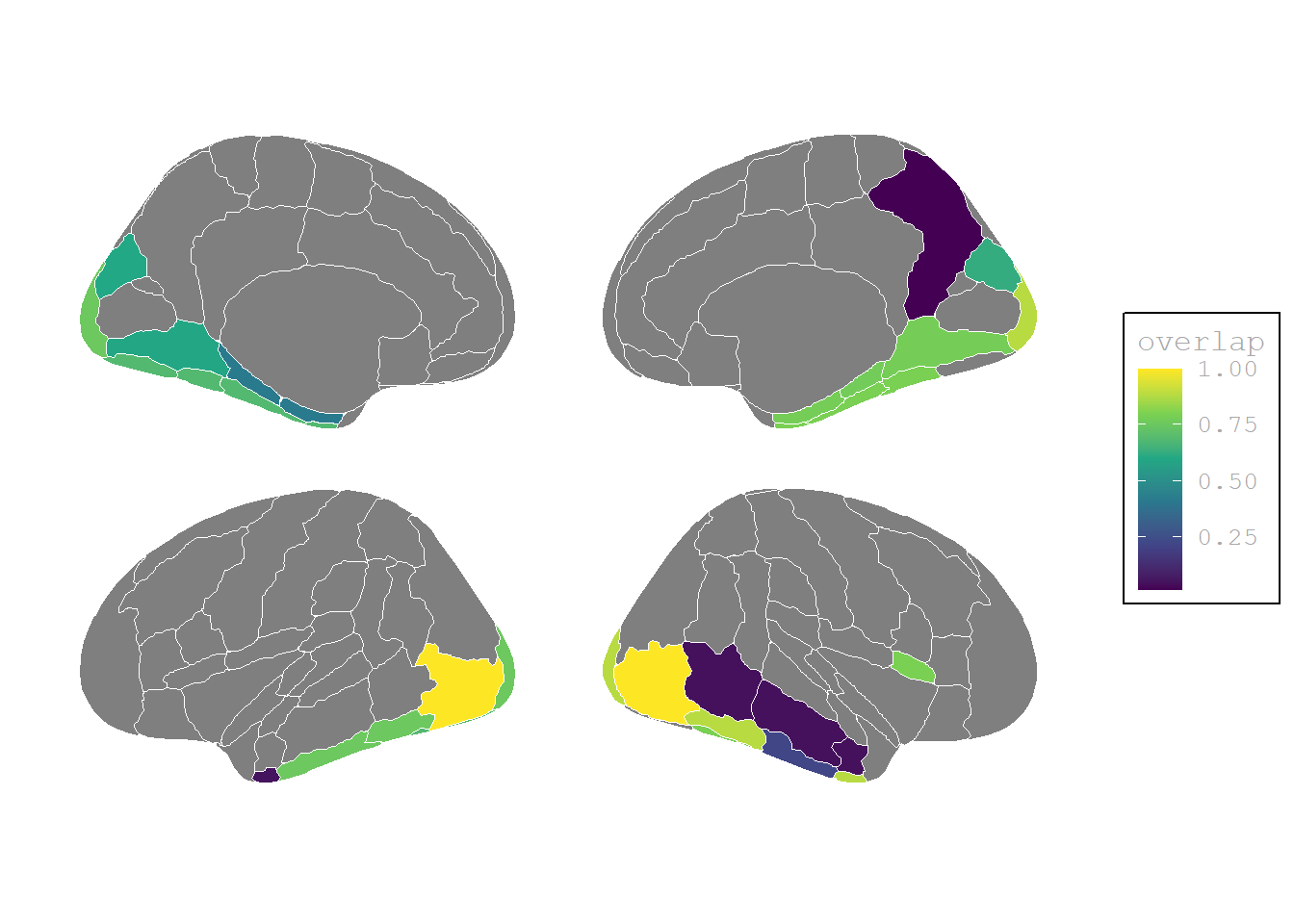
(P<0, I=0, P−I<0)
Code
theme_set(theme_brain2())
tab_4 %>% filter(contrast == "P<0,...,P-I<0") %>%
# left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
left_join(df_regio_abr2, by = c("brain_region" = "reg_abrv")) %>%
rename(hemi = side) %>%
# mutate_at(scale, .vars = vars(C_voxels)) %>%
mutate(overlap = perc_overlap/100) %>%
select(-c(brain_region, # reg_name,
perc_overlap)) %>%
# bind_rows(data.frame(hemi = "right", region = "Temporal Fusiform Cortex ant.", overlap = 1)) %>%
filter(!is.na(region)) %>%
ggplot() +
scale_fill_viridis_c(option = "D", direction = 1) +
geom_brain(atlas = hoCort, # atlas = dk,
color = "white",
position = position_brain(side ~ hemi),
mapping = aes(fill = overlap)) +
labs(fill = "overlap") +
theme(# legend.position = "none",
plot.background = element_blank())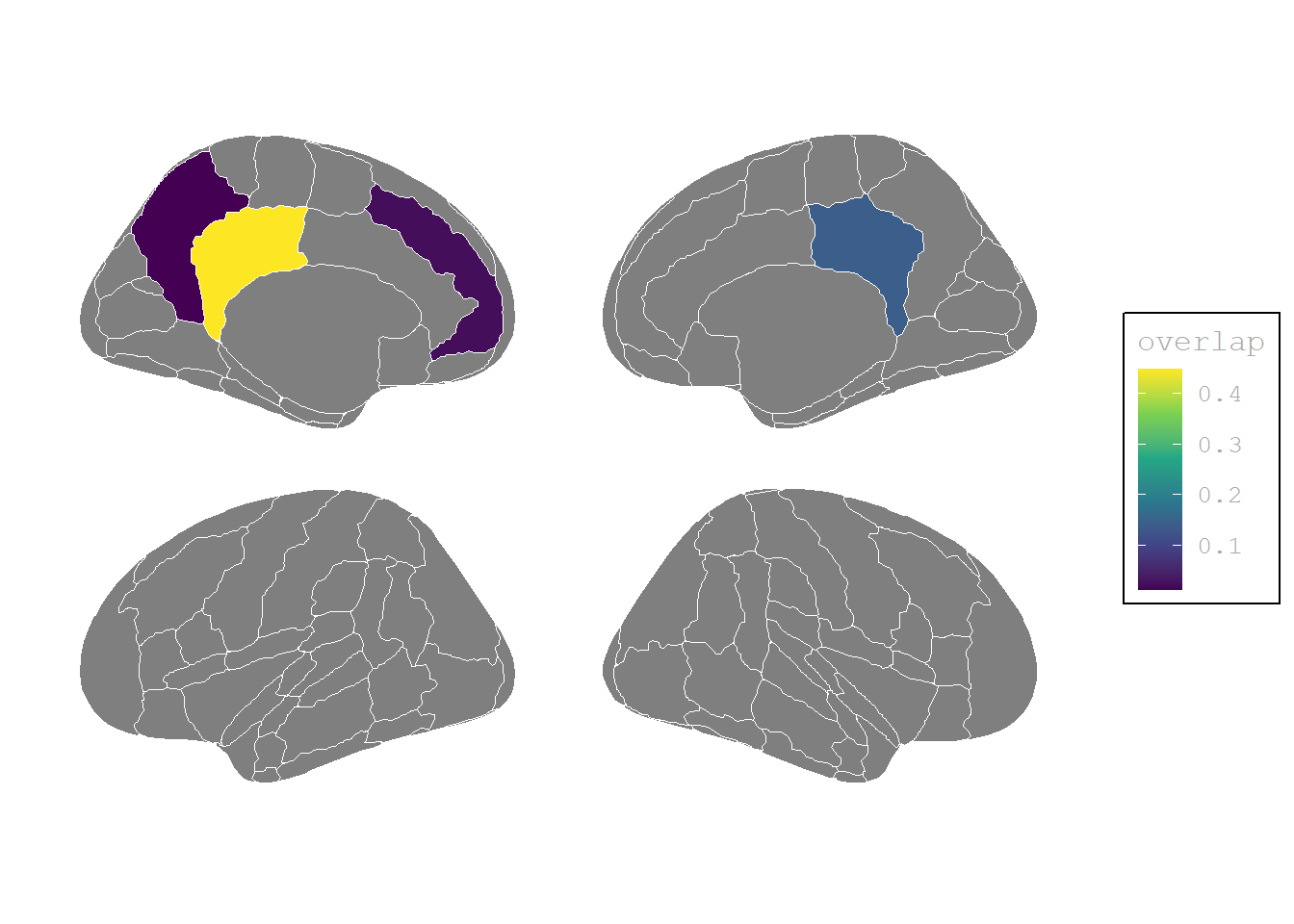
(P>0, I>0, P−I>0)
Code
theme_set(theme_brain2())
tab_4 %>% filter(contrast == "...,I>0,...") %>%
# left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
left_join(df_regio_abr2, by = c("brain_region" = "reg_abrv")) %>%
rename(hemi = side) %>%
# mutate_at(scale, .vars = vars(C_voxels)) %>%
mutate(overlap = perc_overlap/100) %>%
select(-c(brain_region, # reg_name,
perc_overlap)) %>%
# bind_rows(data.frame(hemi = "right", region = "Temporal Fusiform Cortex ant.", overlap = 1)) %>%
filter(!is.na(region)) %>%
ggplot() +
scale_fill_viridis_c(option = "D", direction = 1, begin = 0, end = 1) +
geom_brain(atlas = hoCort, # atlas = dk,
color = "white",
position = position_brain(side ~ hemi),
mapping = aes(fill = overlap)) +
labs(fill = "overlap") +
theme(# legend.position = "none",
plot.background = element_blank())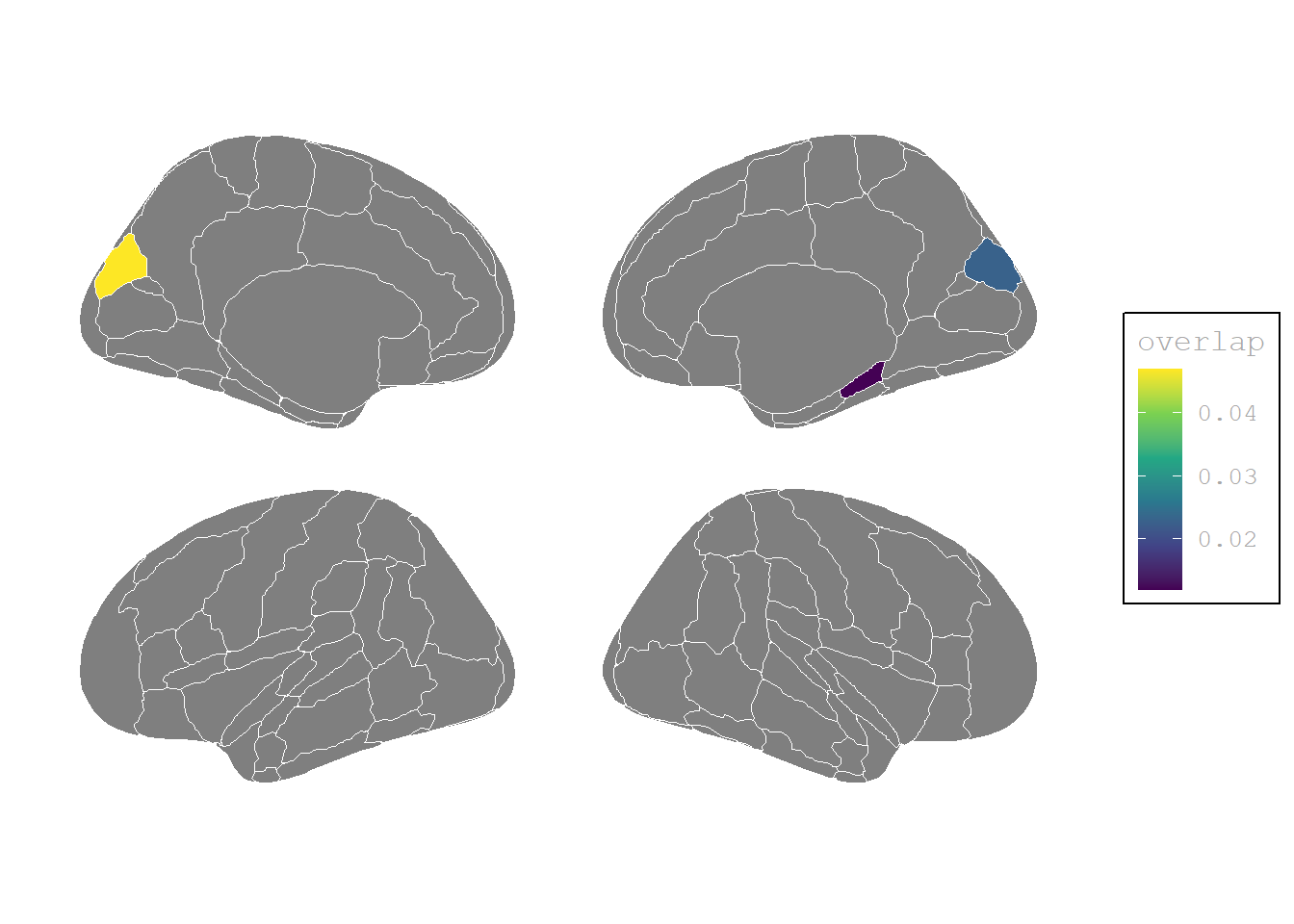
Activation barchart
Code
theme_set(theme_minimal())
# df_regio_abr %>% select(reg_abrv, persp) %>%
# distinct() %>%
# right_join(tab_4, by = c("reg_abrv" = "brain_region")) %>%
# filter(persp != "subcort") %>%
# mutate(rel_overlap = perc_overlap/100) %>%
# # tab_4 %>% mutate(rel_overlap = perc_overlap/100) %>%
# ggplot(aes(x = reorder(reg_abrv, rel_overlap),
# y = rel_overlap, fill = rel_overlap)) +
# geom_bar(stat = "identity") +
# scale_fill_viridis_c(option = "viridis", guide = "none", direction = -1) +
# scale_y_continuous("different voxels", labels = scales::label_percent()) +
# facet_grid(contrast~side, scales = "free", space = "free") +
# # theme(axis.ticks.y = element_blank()) +
# labs(x = "brain region") +
# theme(# text = element_text(size = 20),
# panel.spacing = unit(0.5, "cm")) +
# coord_flip()
# # theme_minimal()
tab_4 %>% mutate(rel_overlap = perc_overlap/100) %>%
ggplot(aes(x = reorder(brain_region, rel_overlap),
y = rel_overlap, fill = rel_overlap)) +
geom_bar(stat = "identity") +
scale_fill_viridis_c(option = "viridis", guide = "none", direction = 1) +
scale_y_continuous("different voxels", labels = scales::label_percent()) +
facet_grid(contrast~side, scales = "free", space = "free", drop = TRUE) +
# theme(axis.ticks.y = element_blank()) +
labs(x = "brain region") +
theme(# text = element_text(size = 20),
panel.spacing = unit(0.5, "cm")) +
coord_flip() -> vxl_chrt_d
vxl_chrt_d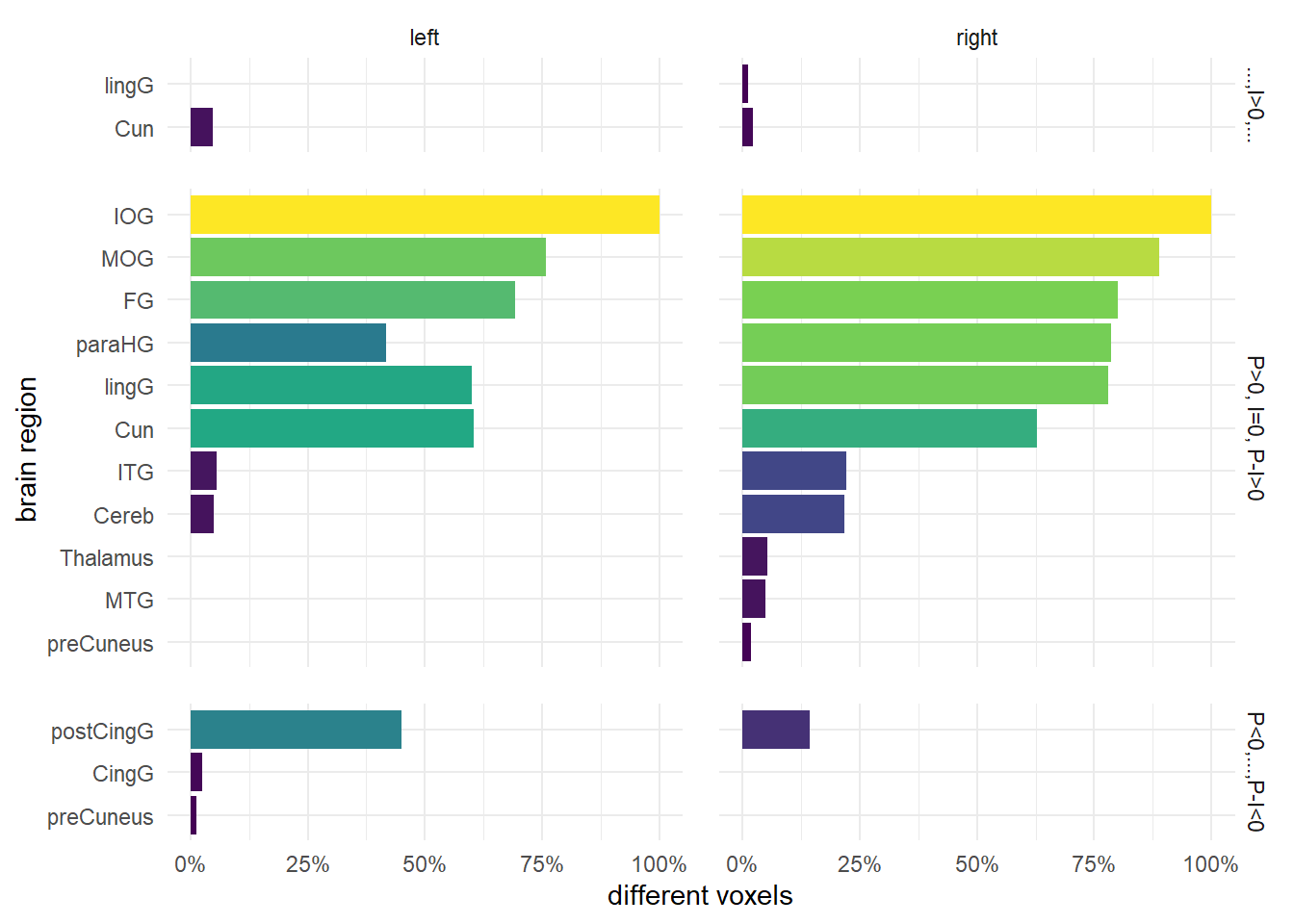
Table D voxels
Code
tab_4 %>% mutate(contrast = factor(contrast,
labels = c("P>0, I>0, P-I>0", "P>0, I=0, P-I>0", "P<0, I=0, P-I<0"))) %>%
pivot_wider(names_from = "side",
values_from = c("perc_overlap", "D_voxels", "x", "y", "z")) %>%
relocate("D_voxels_left", .before = "perc_overlap_left") %>%
relocate("D_voxels_right", .before = "perc_overlap_right") %>%
relocate("y_left", .after = "x_left") %>%
relocate("z_left", .after = "y_left") -> tab_4_wide
df_regio_abr %>% select(c(reg_abrv, reg_name, persp)) %>%
distinct() %>%
right_join(tab_4_wide, by = c("reg_abrv" = "brain_region")) %>%
mutate(brain_region = paste0(reg_name, " (", reg_abrv, ")")) %>%
# filter(persp != "lateral") %>%
select(-c(reg_name, persp, reg_abrv)) %>%
# tab_4_wide %>% left_join(df_regio_abr, by = c("brain_region" = "reg_abrv")) %>%
# mutate(brain_region = paste0(reg_name, " (", brain_region, ")")) %>%
# select(-c(reg_name, persp)) %>%
gt(rowname_col = "brain_region",
groupname_col = "contrast") %>%
tab_stubhead(label = "Contrast") %>%
tab_spanner(label = "left",
columns = c(D_voxels_left, perc_overlap_left, x_left, y_left, z_left)) %>%
tab_spanner(label = "right",
columns = c(D_voxels_right, perc_overlap_right, x_right, y_right, z_right)) %>%
# tab_spanner(label = "left",
# columns = c(x_left, y_left, z_left)) %>%
# tab_spanner(label = "right",
# columns = c(x_right, y_right, z_right)) %>%
cols_label(D_voxels_left = "D voxels",
perc_overlap_left = "%",
D_voxels_right = "D voxels",
perc_overlap_right = "%",
x_left = "x", y_left = "y", z_left = "z",
x_right = "x ", y_right = "y ", z_right = "z ") %>%
cols_align(align = "center", columns = everything()) %>%
sub_missing() %>%
as_raw_html()| Contrast | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| D voxels | % | x | y | z | D voxels | % | x | y | z | |
| P>0, I=0, P-I>0 | ||||||||||
| Inferior temporal gyrus (ITG) | ||||||||||
| Middle temporal gyrus (MTG) | ||||||||||
| Middle occipital gyrus (MOG) | ||||||||||
| Inferior occipital gyrus (IOG) | ||||||||||
| Parahippocampal gyrus (paraHG) | ||||||||||
| Fusiform gyrus (FG) | ||||||||||
| Lingual gyrus (lingG) | ||||||||||
| Precuneus (preCuneus) | ||||||||||
| Cuneus (Cun) | ||||||||||
| Thalamus (Thalamus) | ||||||||||
| Cerebellum (Cereb) | ||||||||||
| P<0, I=0, P-I<0 | ||||||||||
| Posterior cingulate gyrus (postCingG) | ||||||||||
| Precuneus (preCuneus) | ||||||||||
| Cingulate gyrus (CingG) | ||||||||||
| P>0, I>0, P-I>0 | ||||||||||
| Lingual gyrus (lingG) | ||||||||||
| Cuneus (Cun) | ||||||||||